The pipeline of lincRNA identification.: After transcriptome assembly... | Download Scientific Diagram

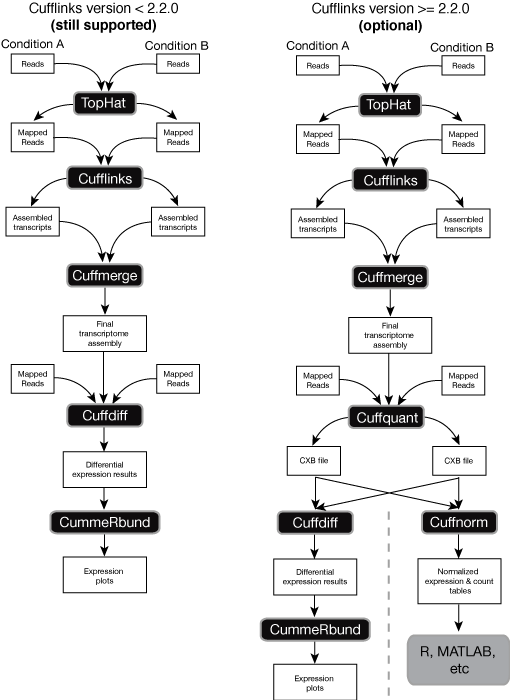
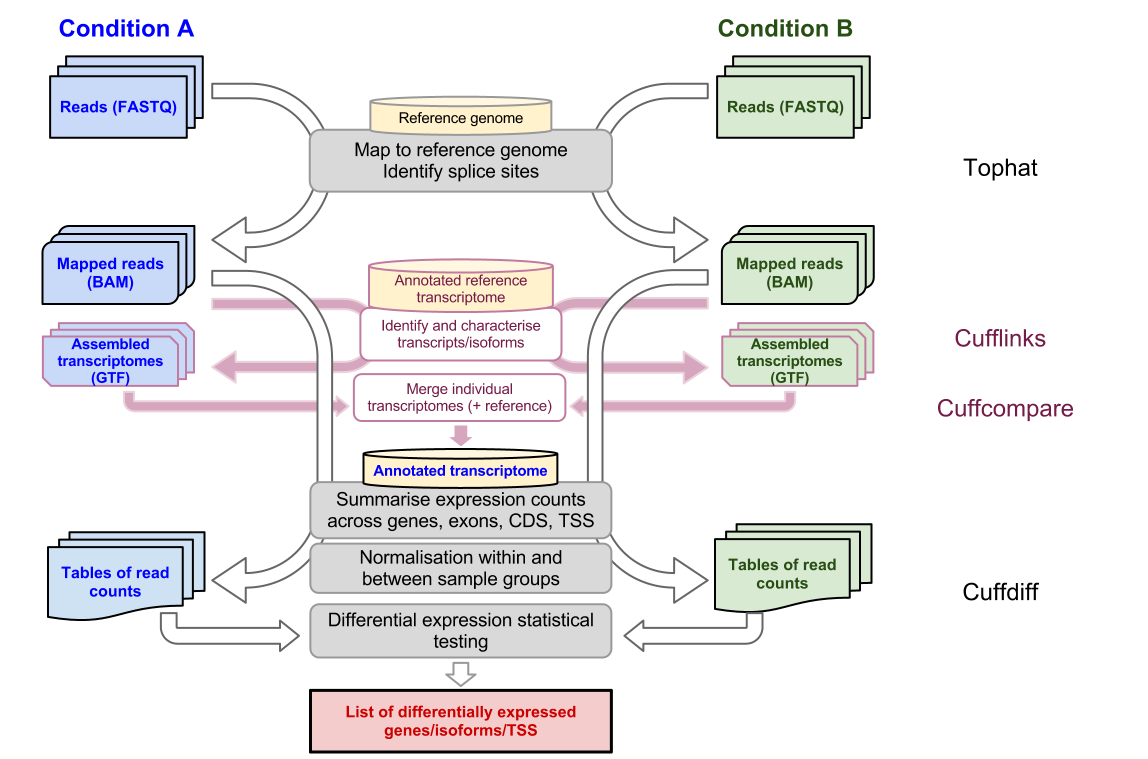
Differential gene and transcript expression analysis of RNA-seq experiments with TopHat and Cufflinks | Nature Protocols
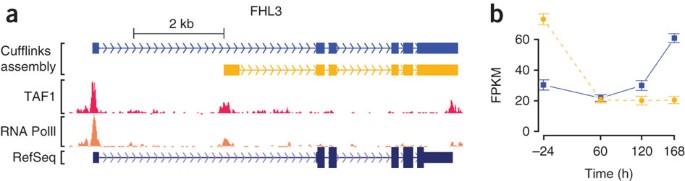
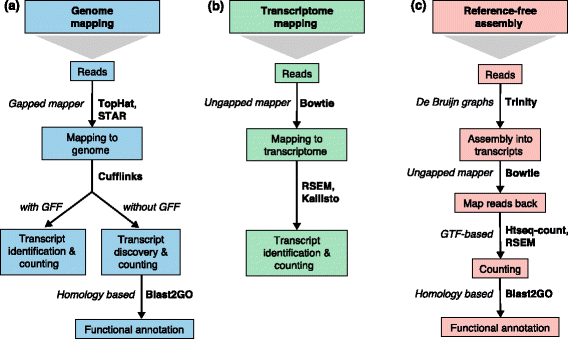
Transcript assembly and quantification by RNA-Seq reveals unannotated transcripts and isoform switching during cell differentiation | Nature Biotechnology

The pipeline. Sixteen transcriptomes were assembled using both de novo... | Download Scientific Diagram
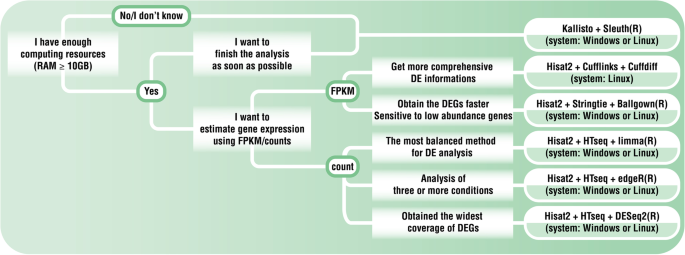
Gene Expression in Eukaryotes | RNA Seq Pipeline | Illumina RNA-Seq | Quality NGS Bioinformatics Data Analysis Services

Establishment of bioinformatics pipeline for deciphering the biological complexities of fragmented sperm transcriptome - ScienceDirect

Transcriptomic Profiling Using Next Generation Sequencing - Advances, Advantages, and Challenges | IntechOpen
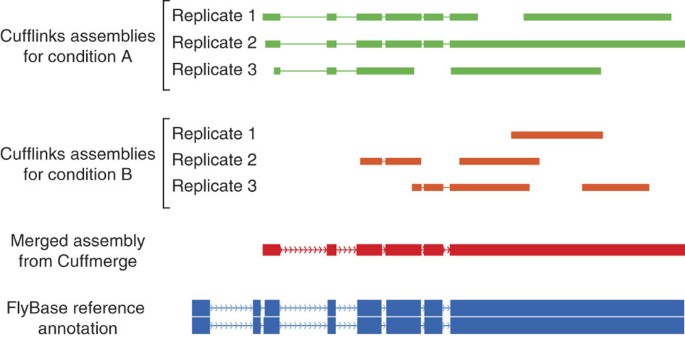
Differential gene and transcript expression analysis of RNA-seq experiments with TopHat and Cufflinks | Nature Protocols

A global survey of the transcriptome of allopolyploid Brassica napus based on single‐molecule long‐read isoform sequencing and Illumina‐based RNA sequencing data - Yao - 2020 - The Plant Journal - Wiley Online Library

Transcript assembly and quantification by RNA-Seq reveals unannotated transcripts and isoform switching during cell differentiation | Nature Biotechnology

Comparison among DiffSplice, FDM, and Cufflinks on simulated data set... | Download Scientific Diagram

Systematic identification and characterization of cardiac long intergenic noncoding RNAs in zebrafish | Scientific Reports

Differential gene and transcript expression analysis of RNA-seq experiments with TopHat and Cufflinks | Nature Protocols